NeutrAvidin beads
NeutrAvidin beads, like Streptavidin beads, can immobilize biotin-labeled compounds and antibodies, DNA, etc. and can be used in a wide range of applications.
NeutrAvidin beads may have slight non-specific adsorption from sugar chains. However, since it does not have a RYD sequence, it suppresses non-specific adsorption derived from cell adhesion proteins compared to Streptavidin beads.
※NeutrAvidin(TM) is a trademark of Thermo Fisher Scientific, Inc. and its subsidiaries.
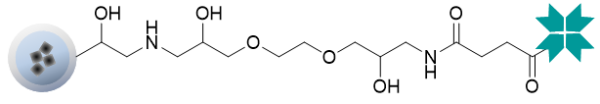

Magnetic beads
The product lineup includes regular FG beads and highly magnetically responsive HM beads.
| Beads | FG beads | HM beads (High Magnetic Response Type) |
|---|---|---|
| Code | TAS8848N1171 | TAB8848N3171 |
| Price | Please contact us | |
| Storage conditions | 2-8℃ (no freezing), protect from light | |
| Storage buffer | 10 mM HEPES(pH7.9), 50 mM KCl, 1 mM EDTA, 10% glycerol |
10 mM HEPES(pH7.9) |
| Magnetization | Superparamagnetism (≧10 emu/g) | Superparamagnetism (≧20 emu/g) |
| Size of beads | 180±30 nm | 140±20 nm |
| Concentration | 20 mg/ml | |
| Functional groups | NeutrAvidin | |
| Binding ability | ≧ 1.0 ug Biotin labeled BSA / mg of beads | |
- Protocol
- SDS
- Papers /
Technical Information - Related Products
- FAQ
Technical Information
-
Cell separation
Learn More→ -
Isolation of drug target protein – Biotinylated MTX
Learn More→
Papers
-
Plakevulin A induces apoptosis and suppresses IL-6-induced STAT3 activation in HL60 cells
Bioorganic & Medicinal Chemistry Letters Volume 110, 15 September 2024, 129886 -
Caspase cleaves Drosophila BubR1 to modulate spindle assembly checkpoint function and lifespan of the organism
The FEBS Journal 290 (2023) 4200–4223 -
Histone deacetylation regulates nucleotide excision repair through an interaction with the XPC protein
iScience 25, 104040, April 15, 2022 -
Telomere-specific chromatin capture using a pyrrole–imidazole polyamide probe for the identification of proteins and non-coding RNAs
Epigenetics & Chromatin (2021) 14:46 -
Directed evolution of orthogonal RNA–RBP pairs through library-vs-library in vitro selection
Nucleic Acids Research, gkab527 (2021) -
Age-related dysfunction of p53-regulated phagocytic activity in macrophages
Biochemical and Biophysical Research Communications 529 (2020) 462 -
The SH3 domain in the fucosyltransferase FUT8 controls FUT8 activity and localization and is essential for core fucosylation
J. Biol. Chem. (2020) 295(23) 7992 -
Tag-Convertible Photocrosslinker with Click-On/Off N-Acylsulfonamide Linkage for Protein Identification
Chemistry—An Asian Journal Volume14, Issue18 September 16, 2019 Pages 3145-3148 -
Chemical Synthesis of Atomically Tailored SUMO E2 Conjugating Enzymes for the Formation of Covalently Linked SUMO–E2–E3 Ligase Ternary Complexes.
J. Am. Chem. Soc., 141, 37, 14742(2019) -
Wisteria floribunda agglutinin staining for the quantitative assessment of cardiac fibrogenic activity in a mouse model of dilated cardiomyopathy
Laboratory Investigation (2019) 99:1749–1765 -
Combinatorially Screened Peptide as Targeted Covalent Binder:Alteration of Bait-Conjugated Peptide to Reactive Modifier
Bioconjugate Chem., 29, 1866 (2018) -
Evaluation of protein-ligand interactions using the luminescent interaction assay FlimPIA with streptavidin-biotin linkage
Analytical Biochemistry 563 (2018) 61-66 -
Generation of Immunity against Pathogens via Single-Domain Antibody–Antigen Constructs
J Immunol December 15, 2016, 197 (12) 4838-4847 -
Alteration of matrix metalloproteinase-3 O-glycan structure as a biomarker for disease activity of rheumatoid arthritis
Arthritis Research & Therapy, 18, 112 (2016). -
Toxic tau oligomer formation blocked by capping of cysteine residues with 1,2-dihydroxybenzene groups
Nature Communications, 6:10216 (2015). -
Tsujita et al.(2013) [ExoCounter]
Ultrahigh-Sensitivity Biomarker Sensing System Based on the Combination of Optical Disc Technologies and Nanobead Technologies
Japanese Journal of Applied Physics 52 (2013) 09LB02 -
A Peptide Derived from Tenascin-C Induces β1 Integrin Activation through Syndecan-4
J. Biol. Chem., 282, 34929 (2007).
- Please tell me how to separate FG beads (magnetic separation and centrifugation).
- Please tell me how to disperse FG beads (ultrasonic method and manual method).
- I mistakenly frozen some beads that were supposed to be stored in the refrigerator. Is it available?
- What amount of the beads is required?
- What are the important points when designing a ligand?
- How are beads stored after the ligands are bound to them?
- Are there any methods other than HPLC for verifying whether or not ligand binding has been successful?
- How strong is the affinity for the proteins that are affinity purified?
- What is the purification efficiency?
- What is the optimal bead type for binding proteins?
- When binding proteins, what should be done if there is lysine residue at a location related to binding with the target substance?
- What is the efficiency when binding proteins?
- What is the optimal bead type for binding peptides?
- How is the cell extract prepared?
- Is there any problem with using frozen stock homogenate?
- How much protein supply is necessary?
- Can affinity purification be used with membrane proteins such as GPCRs and ion channels?
- There are many background bands. how can i reduce it?
- What should be done when a large number of bound protein bands are detected?
- Is it necessary to use the recommended buffer as the binding buffer?
- Why is it that both salt elution and boil elution are performed for elution?
- Does it happen that the band of bound protein becomes thin when the concentration of ligand is increased?
- Why can’t I see any bands of bound proteins?
- How long is the stable period of the ligand-immobilized beads?
- Is the optimal binding reaction time of 4 hours?
- What is the optimal bead type for immobiliding antibodies?
- Can I quantify the immobilization amount of the antibodies on the beads?
- What is the efficiency when immobiliding antibodies?
- Is there a way to increase the antibody immobilization efficiency (immobilization amount)?
- Can I disperse antibody-immobilized beads by ultrasonic device?
- When immobilizing antibodies to beads, the beads may adhere to the wall of the tube. Is there a way to suppress this?
- How can I improve dispersibility of antibodies immobilized beads?
- I want to analyze bound proteins with MS, but what should I do if the target protein band is thin?
- How much protein can be analyzed by MS?

