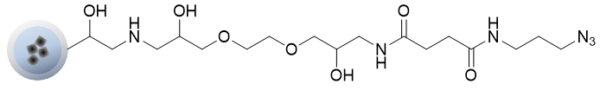
Azide beads
Azide beads can be easily immobilized on beads by a click chemistry reaction of compounds with alkynes.
In the case of a compound having multiple immobilizable sites such as an amino group, the compound can be immobilized on the beads in a site-specific manner by introducing an alkyne to the site to be immobilized.

Magnetic beads
The lineup of this product is only regular FG beads.
| Beads | FG beads |
|---|---|
| Code | TAS8848N1160 |
| Price | Please contact us |
| Storage conditions | 2-8 ℃ (no freezing) |
| Storage buffer | Ultrapure water |
| Magnetization | Superparamagnetism (≧10 emu/g) |
| Size of beads | 180±30 nm |
| Concentration | 20 mg/ml |
| Functional groups | Azide group |
| Amounts of the functional groups | 70-140 nmol/mg of beads |
- Protocol
- SDS
- Papers /
Technical Information - Related Products
- FAQ
- Screening by using ligand immobilized beads
- Competitive inhibition
- Drug elution
- Immobilization of ligands (alkyne structure compounds) on azide beads using click chemistry reaction
- Immobilization of ligands (alkyne structure compounds) on azide beads using click chemistry reaction (Small scale method)
- Preparation of cell extract (Large scale method)
- Preparation of cell extract (Small scale method)
Technical Information
Papers
-
Identification of a novel target of sulforaphane: Sulforaphane binds to acyl-protein thioesterase 2 (APT2) and attenuates its palmitoylation
Biochemical and Biophysical Research Communications 726 (2024) 150244 -
Novel ability of diflubenzuron as an inhibitor of mitochondrial function
Insect Biochemistry and Molecular Biology Volume 167, April 2024, 104088 -
Sulforaphane suppresses the activity of sterol regulatory element-binding proteins (SREBPs) by promoting SREBP precursor degradation
Scientifc Reports (2022) 12:8715 -
Dioctatin Activates ClpP to Degrade Mitochondrial Components and Inhibits Aflatoxin Production
Cell Chemical Biology 27 (2020) 1396 -
An acyl-SAM analog as an affinity ligand for identifying quorum sensing signal synthases
Chem. Commun., DOI:10.1039 (2014). -
Protein fishing using magnetic nano-beads containing calmodulin site-specifically immobilized via an azido-group
J. Biochem., DOI:10.1093 (2013).
- Please tell me how to separate FG beads (magnetic separation and centrifugation).
- Please tell me how to disperse FG beads (ultrasonic method and manual method).
- I mistakenly frozen some beads that were supposed to be stored in the refrigerator. Is it available?
- What amount of the beads is required?
- What are the important points when designing a ligand?
- How are beads stored after the ligands are bound to them?
- Are there any methods other than HPLC for verifying whether or not ligand binding has been successful?
- How strong is the affinity for the proteins that are affinity purified?
- What is the purification efficiency?
- How is the cell extract prepared?
- Is there any problem with using frozen stock homogenate?
- How much protein supply is necessary?
- Can affinity purification be used with membrane proteins such as GPCRs and ion channels?
- There are many background bands. how can i reduce it?
- What should be done when a large number of bound protein bands are detected?
- Is it necessary to use the recommended buffer as the binding buffer?
- Why is it that both salt elution and boil elution are performed for elution?
- Does it happen that the band of bound protein becomes thin when the concentration of ligand is increased?
- Why can’t I see any bands of bound proteins?
- How long is the stable period of the ligand-immobilized beads?
- Is the optimal binding reaction time of 4 hours?
- I want to analyze bound proteins with MS, but what should I do if the target protein band is thin?
- How much protein can be analyzed by MS?

